 
Both p K a and isoelectric point estimates have been used in numerous techniques, such as two-dimensional gel electrophoresis (2D-PAGE) ( 3, 4), crystallization ( 5), capillary isoelectric focussing ( 6), and mass spectrometry (MS) ( 7, 8). Taken together, the p K a values of all charged groups can be used to calculate the overall charge of the molecule in any pH or to estimate the isoelectric point ( pI, IEP), that is, the pH at which there is an equilibrium of positive and negative charges and therefore the total net charge of the molecule is equal to zero ( 2). For proteins and peptides, the ionizable groups of seven charged amino acids should be considered: glutamate (γ-carboxyl group), cysteine (thiol group), aspartate (β-carboxyl group), tyrosine (phenol group), lysine (ε-ammonium group), histidine (imidazole side chains), and arginine (guanidinium group) ( 1). The charge of a protein is one of its key physicochemical characteristics and is related to the p K a dissociation constant (p K a is a quantitative measure of the strength of an acid in solution). In addition, isoelectric points for proteins in NCBI non-redundant (nr), UniProt, SwissProt, and Protein Data Bank are available in both CSV and FASTA formats. The database enables the retrieval of virtual 2D-PAGE plots and customized fractions of a proteome based on the isoelectric point and molecular weight. To facilitate bottom-up proteomics analysis, individual proteomes were digested in silico with the five most commonly used proteases (trypsin, chymotrypsin, trypsin + LysC, LysN, ArgC), and the peptides’ isoelectric point and molecular weights were calculated. The isoelectric point for proteins is predicted by 21 methods, whereas pKa values are inferred by one method. The Proteome- pI 2.0 database includes data for over 61 million protein sequences from 20 115 proteomes (three to four times more than the previous release). Additionally, it can be obtained directly from the p K a values of individual charged residues of the protein. The isoelectric point-the pH at which a particular molecule carries no net electrical charge-is an important parameter for many analytical biochemistry and proteomics techniques. To assist in determining similarity we define two classes of acids.Proteome- pI 2.0 is an update of an online database containing predicted isoelectric points and p K a dissociation constants of proteins and peptides. If additional acidic or basic groups are present as side-chain functions, the pI is the average of the pK a's of the two most similar acids. Thus, the pI for alanine is calculated to be: (2.34 + 9.69)/2 = 6.02, the experimentally determined value. For simple amino acids such as alanine, the pI is an average of the pK a's of the carboxyl (2.34) and ammonium (9.69) groups. In other words, the positively charged groups are exactly balanced by the negatively charged groups. 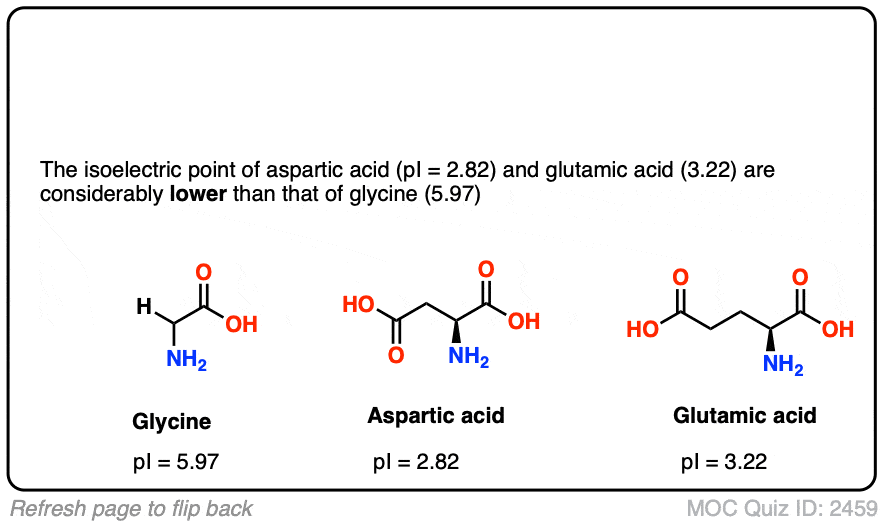
The isoelectric point, pI, is the pH of an aqueous solution of an amino acid (or peptide) at which the molecules on average have no net charge. The very high pH required to remove the last acidic proton from arginine reflects the exceptionally high basicity of the guanidine moiety at the end of the side chain. Formulas for these species are written to the right of the titration curves, together with the pH at which each is expected to predominate. For each of these compounds four possible charged species are possible, one of which has no overall charge. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
